 |
 |
- Search
| Healthc Inform Res > Volume 27(4); 2021 > Article |
|
Abstract
Objectives
Different complex strategies of fusing handcrafted descriptors and features from convolutional neural network (CNN) models have been studied, mainly for two-class Papanicolaou (Pap) smear image classification. This paper explores a simplified system using combined binary coding for a five-class version of this problem.
Methods
This system extracted features from transfer learning of AlexNet, VGG19, and ResNet50 networks before reducing this problem into multiple binary sub-problems using error-correcting coding. The learners were trained using the support vector machine (SVM) method. The outputs of these classifiers were combined and compared to the true class codes for the final prediction.
Results
Despite the superior performance of VGG19-SVM, with mean ± standard deviation accuracy and sensitivity of 80.68% ± 2.00% and 80.86% ± 0.45%, respectively, this model required a long training time. There were also false-negative cases using both the VGGNet-SVM and ResNet-SVM models. AlexNet-SVM was more efficient in terms of running speed and prediction consistency. Our findings also showed good diagnostic ability, with an area under the curve of approximately 0.95. Further investigation also showed good agreement between our research outcomes and that of the state-of-the-art methods, with specificity ranging from 93% to 100%.
Conclusions
We believe that the AlexNet-SVM model can be conveniently applied for clinical use. Further research could include the implementation of an optimization algorithm for hyperparameter tuning, as well as an appropriate selection of experimental design to improve the efficiency of Pap smear image classification.
Cancer refers to the uncontrollable multiplication and growth of cells that are either localized (carcinoma in situ [CIS]) or have spread to other parts of the body. Cervical cancer, although treatable when detected early, is the most common form of cancer in women aged 35 and younger [1]. The prevalence of cervical cancer is disproportionately high in individuals who have sexual relationships at early ages, who have sexually transmitted infections, and who use tobacco [1,2]. Therefore, cervical screening is highly recommended for women aged between 21 and 65 years once every 3 years [3,4]. However, as many people have skipped screening appointments and avoided visiting healthcare facilities due to infection-related fears amid the recent coronavirus disease 2019 pandemic [5], concerns have grown regarding the possible demand for healthcare resources once the pandemic is over [6].
There is a gradual progression leading to cervical cancer, from normal squamous (NS), to dysplasia (or precancerous cells), and finally to invasive disease. Cytological screening for cervical cell abnormalities using the Papanicolaou (Pap) smear is an effective method to detect precancerous cells. These precancerous stages (or cervical intraepithelial neoplasia [CIN]) are classified as (1) CIN I or low-grade dysplasia (LGD), (2) CIN II or high-grade dysplasia (HGD), or (3) CIN III or CIS, depending on the extent to which squamous cells fail to mature as they migrate to the surface of the epithelium [7]. Dysplastic cells have nuclei that occupy a large fraction of the intracellular compartment and display nuclear membrane irregularities. There is also an increase in the nuclear-to-cytoplasmic ratio (NCR) from 1:7 in normal squamous cells to 1:1.5 in the CIN III group [8]. Therefore, the cell features used by cyto-technicians in the diagnosis include aspects of cell appearance such as size, color (after staining), textures and shapes, NCR, and the distribution of nuclei [9]. However, not only is this approach subjective and prone to inter- and intra-observer variability, it requires experience and a trained eye, and is time-consuming.
As internet-accessible databases continue to grow, accompanied by advances in computing techniques (e.g., parallel programming and edge computing), these resources have contributed to major milestones in artificial intelligence research. At present, most of the computer-aided diagnostic (CAD) systems used for cervical cancer detection require the extraction of engineered features, such as shape-based features [10,11], morphological features [11], and statistical features [12], used by a classifier in its diagnosis. Classifiers such as decision forests, artificial neural networks, K-nearest neighbor models, and support vector machines (SVM) have been used with considerable success for binary or multiclass problems in various domains. Among these classifiers, SVM has received relatively more research attention and has been adopted in various areas [13–15]. This model has also been tested for its superiority in cervical cell recognition [16,17]. This superiority is attributed to its robustness to noise and outliers, its high susceptibility to overfitting, and its capability to handle nonlinearly separable data.
There is growing progress in the use of pretrained deep networks such as LeNet [18], Inception-ResNet [19], Visual Geometry Group network (VGGNet) [20], ShallowNet [21], and deep ConvNet [22], which can be applied to cervical cell grading through transfer learning. These models were reported to provide a higher classification accuracy than training a neural network from scratch [23,24]. At present, most research has studied only two-class problems: normal and abnormal [18,21,22]. Those studies have reported accuracies nearing 100% using a hybrid of deep learning and handcrafted features. A major effort that is worth mentioning is the fusion technique highlighted by Jia et al. [18]. Their study demonstrated that a combination of features from various layers of a convolutional neural network (CNN) and strong features (textural, morphological, and chromatic) extracted from cervical cell images enhanced prediction accuracy (up to 99%) using an SVM-based model. A similar approach was adopted in another study [25] that observed a lower average accuracy of approximately 80% using whole slide images. This method of feature fusion can be a tedious and time-consuming process, as it involves the laborious task of manual optimization and iterative feature engineering.
Since different treatment regimens should be used for lesions of different dysplasia categories [26,27], a CAD system able to diagnose multiple lesion types would enable clinicians to establish the most appropriate therapeutic strategies and prevent the risk of progression to advanced stages. For this reason, researchers [24] extended early efforts [17,19,23] and studied up to four-class problems with NS, LGD, HGD, and CIS. We sought to expand this line of research by including cervical columnar cell types as an additional condition. This type of cell has borderline risk (i.e., it is neither normal nor with evidence of dysplasia [28]), so it is placed in a separate class of problems. The main contributions of this paper include: (1) a simplification of earlier attempts using binary partitions of extracted deep features for the classification task, (2) a five-class cervical cell image classification using joint CNNs and an error-correcting SVM method, and (3) a discussion of key factors affecting the performance of classification systems and future improvement opportunities. This work was carried out using MATLAB 2020a.
Below are presented brief descriptions of the images used and the image processing steps carried out in this study. Sections II-2 and II-3 deal with the methodologies used in our work.
We carried out our research on a benchmark dataset containing 917 single cervical-cell images downloaded from Herlev University (http://mde-lab.aegean.gr). This is, to our best knowledge, the only public database that contains a comprehensive number of cervical cell types. This data source allowed us to assess the performance of our models relative to the task requirements and facilitated comparison with previous research that used the same dataset [16,18–22]. This dataset comprises seven labeled classes of cervical cells with clinical outcomes commonly examined in cervical cancer research: normal (i.e., superficial, intermediate, and columnar squamous) and abnormal (i.e., mild, moderate, and severe dysplasia, and CIS). These images are grouped, in our paper, into the following five classes, as discussed above: NS, columnar squamous (CS), LGD, HGD, and CIS (Figure 1). Polygonal-shaped superficial and intermediate cells represent normal cervical epithelial cells (NS) [19]. They are found in abundance in the superficial and intermediate layers of the squamous epithelium, and their relative numbers vary depending on the stage of the ovulation cycle. Cases of mild dysplasia are classified as LGD, while cases of moderate and severe dysplasia are grouped under HGD in accordance with the Bethesda system. These classes had an imbalanced data distribution, as shown in Table 1. They were randomly split using a random seed number of 1 and a data partition ratio of 0.6:0.15:0.25 for training, validation, and model testing, as shown in Figure 2. The training data size was enlarged using an augmentation strategy to prevent overfitting. This was done by rotating the original training images by 10°, −10°, 20°, −20° angles and vertically flipping the images. These color images were resized to 227 × 227 × 3 and 224 × 224 × 3 according to the acceptable input size for the different network models used in this work.
The deep CNN models used in our experiment for supervised learning were AlexNet, VGG19, and ResNet50 due to the simplicity (in the case of AlexNet) and uniformity (i.e., VGGNet) of their architecture. ResNet is a stack of residual blocks that solves the vanishing gradient problems with skipping connections (dashed lines) for better training.
We first trained the pretrained CNN models with all layers frozen for the cervical cell classification task with the training dataset (including augmented images) in Table 1. The softmax loss function was used as the activation function in this end-to-end training process (as shown at the center of Figure 2). These models were tested and fine-tuned using a validation set that was not part of the training set to provide evidence of over- or under-fitting of the data during each training session. The final training was performed with the sdgm optimizer, 30 training epochs (N), a mini batch size (β) of 32, and a gradient threshold (ψ) and initial learning rate (η) of 0.01 and 0.008, respectively. This was the best hyper-parameter set determined from a fixed grid search space. In the experiment, 35 iterations were performed to test different parameter sets randomly combined from: optimizer: {adam, sgdm}, N = {10, 20, 30}, β = {20, 32, 64}, ψ = {0.01, 0.1, 1}, and η = {0.001, 0.005, 0.008, 0.01, 0.05, 0.1}. We then extracted the learned features from the first fully connected (fc) layer in each trained model for further training using the SVM classifier system to enhance the model’s classification performance. The code was run on an Intel Xeon E52680. Each model was consecutively trained three times and the average run time was recorded.
The SVM method is based on the construction of single or multiple hyperplanes to solve nonlinear problems via matching attributes of observations to a class label. The SVM technique used in this research was combined with the error-correcting output codes (ECOC) model (using the fitcecoc function in MATLAB) to decompose a multiclass classification problem into several binary ones. Although some of the algorithm’s parameters can be configured by users to optimize the classification performance, the default settings were used in this study. The ECOC method uses (k(k–1))/2 learners for k classes. Each binary learner is trained on feature vectors of the training data of two class labels, as shown in Table 2, using SVM. For example, in the case of learner 1, all observations in the CIS class were grouped into a positive class (+1), observations of HGD into a negative class (−1), and those that belonged to other classes were ignored before this learner is trained on them. This is repeated for the remaining learners. During the classification of a new (unseen) observation, the outputs of the classifiers are combined to form an output code. This code was compared to the 10-bit codes in Table 2. The difference was optimized using the aggregated losses of the quadratic function, L, given in Equation (1) [29]. The class with the nearest code to the output code was assigned as the class label.
where n is the sample size, w
j
is the weight for observation j, y
j
is the corresponding class label and f(X
j
) is the classification score for observation j of the predictor data X. We used this multi-learner classifier to complement the softmax function in the original network, which was reported to be ineffective at minimizing intra-subject variation [30], and to enhance the decision regarding the diagnosis. This classifier system used the features (from fc6 in AlexNet and VGG, and from fc in ResNet) in this second-stage training of the CNNs. The results of our experiment showed a higher mean prediction accuracy (approximately 4%) using this technique than using softmax alone. The largest increases were recorded as 2%, 4%, and 9%, respectively, using the trained AlexNet, ResNet-50 and VGG19, and SVM jointly. In this paper, only results from the SVM classifier are reported in detail.
The classification performance of CNN-SVM models used in this work was compared using the mean accuracy, specificity, and recall or sensitivity measures given in Equations (2)–(4). While accuracy describes the closeness of the prediction to the ground truth, both specificity and sensitivity quantify the efficiency of a model in differentiating the classes of cervical cells. Table 3 shows our models’ performance evaluated on the testing dataset and average simulation run time for three consecutive training sessions.
where M denotes the total class labels (M = 5). A true positive (TP
i
) is a class member correctly predicted as a specific class label, i. A false positive (FP) is a nonclass member incorrectly predicted as a class member, a false negative (FN) is a class member misclassified as a nonclass member, and a true negative (TN) is a nonclass member correctly classified as a nonclass member.
Before moving on to present the results from each model in Table 3, it is worth mentioning that we noticed cases of false negatives (testing images of other classes were being misclassified as NS) by VGG19-SVM and ResNet50-SVM in every run (i.e., a non-repeated image from the 226 images in each run), but no such cases were seen using AlexNet-SVM. Highly similar values of the evaluated performance metrics between the models can also be observed from this table. Therefore, the best-performing models (shown in bold in Table 3) were further investigated and discussed. Their confusion matrix and receiving operating characteristic (ROC) curves (i.e., performance graphs) from a one versus all analysis are plotted in Figure 3. The performance comparison based on sensitivity and specificity measures in Figure 4 showed the same adequacy of the models in detecting the NS class label, while the poorest performance was seen in the HGD categories. The area under the curve (AUC) for each diagnosis class showed values ranging from 0.85 to 1, as shown in Figure 4, indicating the overall high reliability of our diagnostic system in differentiating normal from abnormal cells and in the grading of dysplastic images. A comparison of the performance of our method with that of other recently reported approaches is shown in Table 4.
In this study, we adopted different transfer-learned CNNs combined with an SVM classifier system for the classification of cervical Pap smear images. Our findings, shown in Table 3, suggest good consistency in both intra- and inter-model comparisons of classification performance. The ResNet slightly underperformed as compared to the more complex VGGNet, suggesting that the recovery of the lost information (in ResNet) in the training phase may not significantly improve the accuracy.
Even though all the models used in the present study accurately identified normal cervical cases (with excellent specificity and sensitivity of 100%, and an AUC of 1), the VGGNet and ResNet models exhibited some cases where cervical cell images of other classes were misclassified as normal. An investigation of these images showed that they had overlapping cells, poor contrast, and cropped cells at image borders. These FN results are a matter of concern, as the patient would miss the optimum time for treatment that would maximize therapeutic outcomes. Meanwhile, the AlexNet-SVM model demonstrated good estimation performance, in terms of high sensitivity and specificity, good reproducibility, and fast training speed, as shown in Table 3. We observed a similar classification performance between AlexNet-SVM and VGG19-SVM (overall accuracy >80%; mean AUC = 0.95), with the former requiring only a fraction of the computational time in the training (by an order of magnitude) than other models. Both models demonstrated good performance in the classification of CIS and HGD (i.e., specificity and sensitivity ≥80%), as shown in Figure 4, but they had considerably inferior sensitivity performance (approximately 60%–70%) in the NC and LGD classes. Our results showed that several NC images (approximately 25%) were misclassified as HGD, as shown in Figure 3. This resulted in high FP rates (about 20%) in HGD and lower AUC values (0.86–0.89) than with other class labels. Similarly, about one-third of LGD images were misclassified as NC and HGD. This likely occurred because of gradual and minute changes in the characteristics of these cervical cell images (e.g., nuclei size, NCR) as the lesion progressed, as shown in Figure 1. Some features important for the diagnosis may not have been sufficiently and consistently captured in the employed dataset. For example, some missing cell borders and variability in the nuclei size ranges (possibly due to inconsistent cropping) were identified in most of the images used. The cell characteristics in the current dataset exhibit subtle and complex differences, which can be difficult to discern using the current architectures. Instead, irrelevant information may have passed through the networks. This was aggravated by the increased number of considered class labels, which reduced the overall classification performance, as has also been reported in previous research [20,25]. This may prompt a search for an enhanced network architecture, such as the use of extra convolutional or pooling blocks to preserve more relevant features from primary feature maps in the near future. The classification performance may also be enhanced by using a dataset with consistent imaging conditions (e.g. brightness, exposure/illumination, and viewing conditions) and fine-tuning of hyperparameters, which can make the process less laborious, while also covering a larger search space using an optimization algorithm.
Most of the recently reported studies in Table 4 have recommended introducing either adaptive feature selection or fusion into deep neural network models to enhance classification performance. However, this strategy can be experimentally exhaustive and time-consuming. Our research simplifies the earlier designs and focuses on using deep features of trained CNNs and the SVM model for classification tasks. The comparison between methods in Table 4 shows that the architecture and use of CNNs have a significant influence on classification performance. From this table, we identified the experimental group and design as the two key factors affecting prediction accuracy. The closest work to ours, which considered up to four-class problems [24], observed a higher accuracy (>90%), which can be attributed to the use of a larger and independent dataset.
In conclusion, based on our analyses, AlexNet-SVM is preferable for this task. This model can be easily and conveniently extrapolated to clinical practice to support the diagnoses made by professionals. Our system has comparable efficacy to that of state-of-the-art systems in identifying normal cells, thereby eliminating unnecessary biopsies to confirm the diagnosis. Meanwhile, more precise dysplasia grading may be achieved with the use of a meticulously designed network and by including a larger dataset with consistent imaging conditions in the training process.
Acknowledgments
This research was supported by Ministry of Higher Education Malaysia (MOHE) through Fundamental Research Grant Scheme (No. FRGS/1/2020/TK0/UTHM/02/27). We also want to thank Universiti Tun Hussein Onn Malaysia for partially funding this work (No. TIER1-H766).
Figure 1
Cervical cell classes: (A) normal squamous, (B) normal columnar, and (C) low-grade dysplasia; (D) high-grade dysplasia (HGD) with moderate dysplasia, (E) HGD with severe dysplasia, and (F) carcinoma in situ.
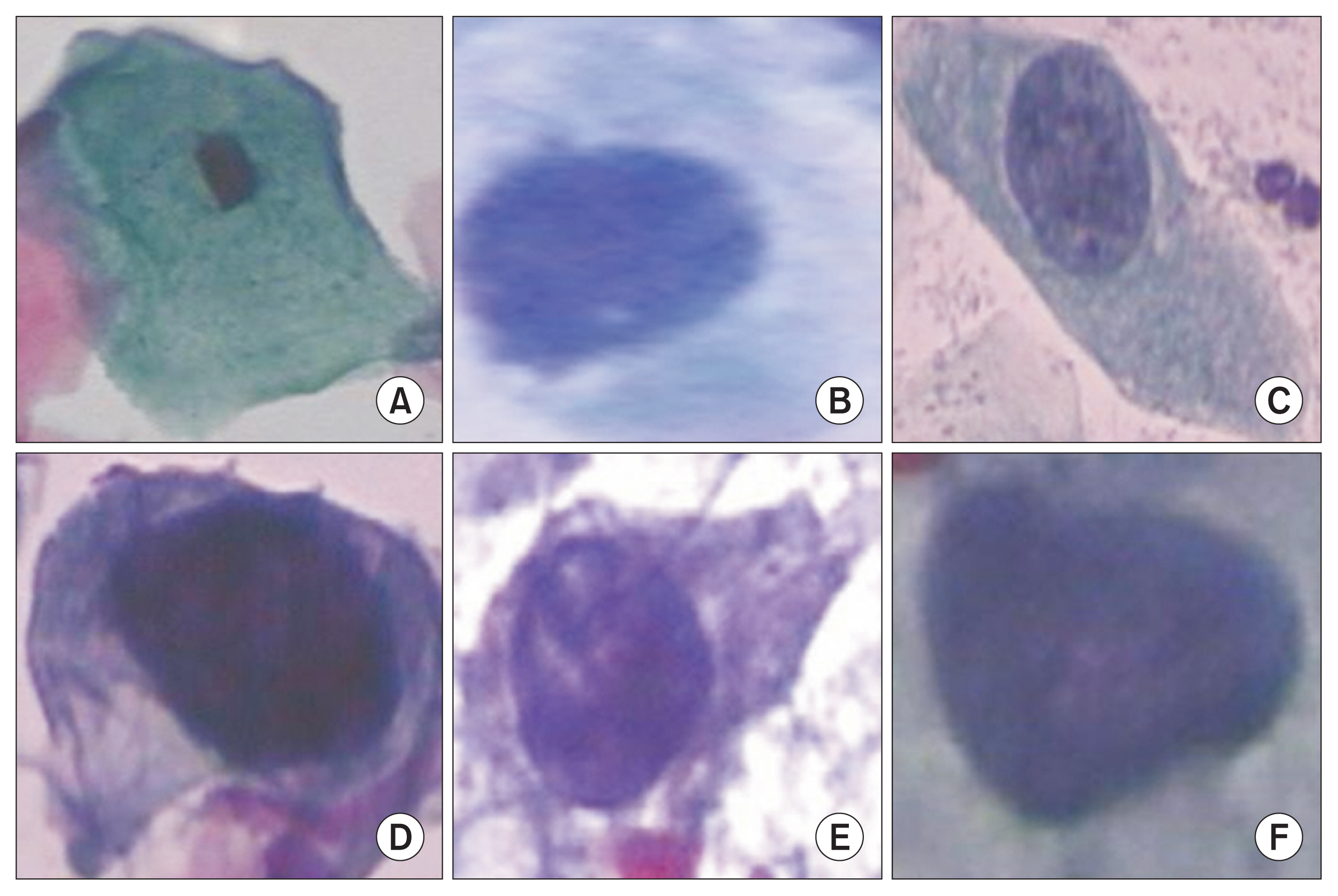
Figure 2
Joint CNN-SVM framework and the classification pipeline. CNN: convolutional neural network, SVM: support vector machine, fc: fully connected, CIS: carcinoma, HGD: high grade, LGD: low grade dysplasia, NC: normal columnar, NS: normal squamous.
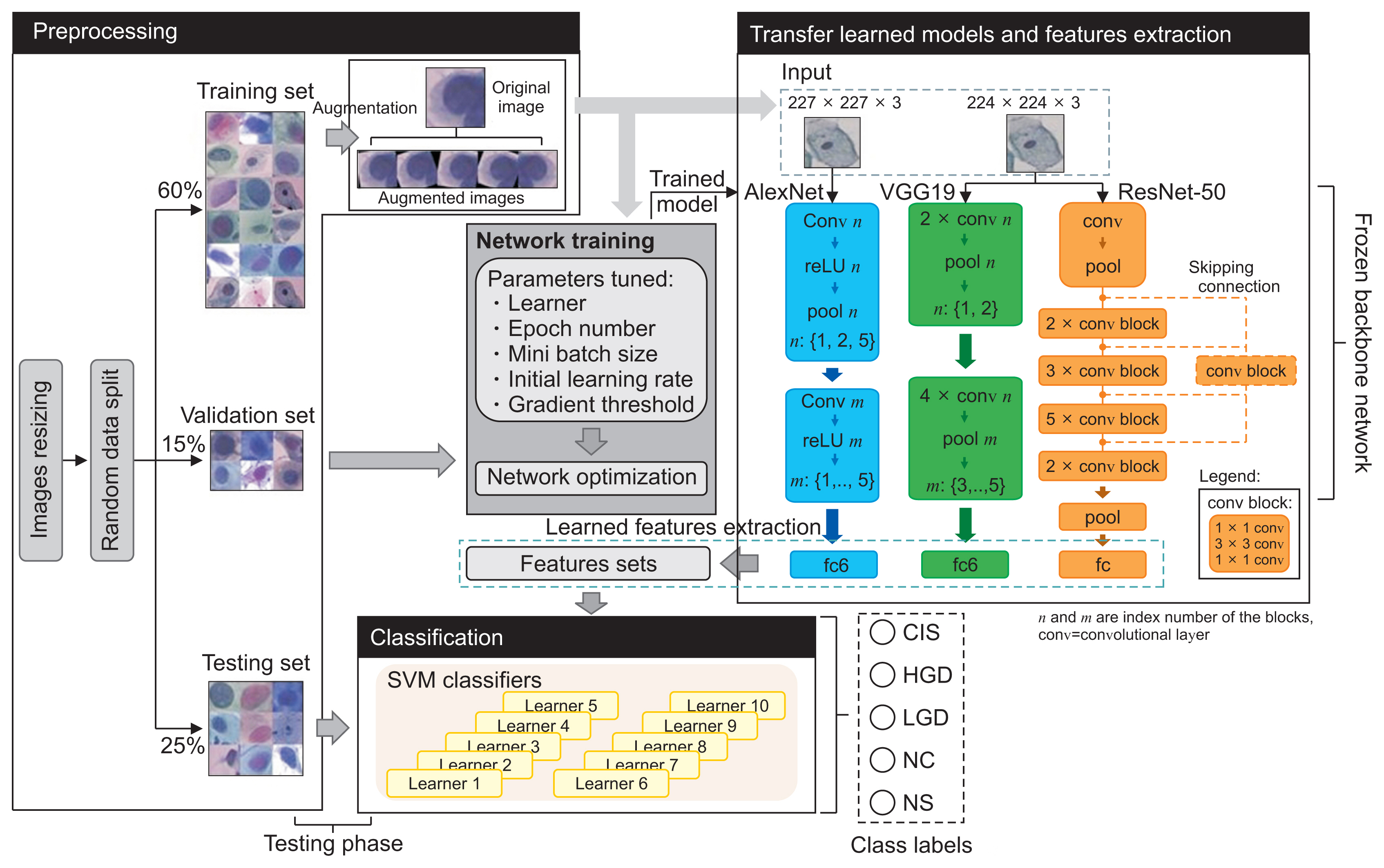
Figure 3
Confusion matrices and performance curves of the best-performing models: (A) AlexNet-SVM, (B) VGG19-SVM, and (C) ResNet50-SVM. CIS: carcinoma, HGD: high grade, LGD: low grade dysplasia, NC: normal columnar, NS: normal squamous, SVM: support vector machine.
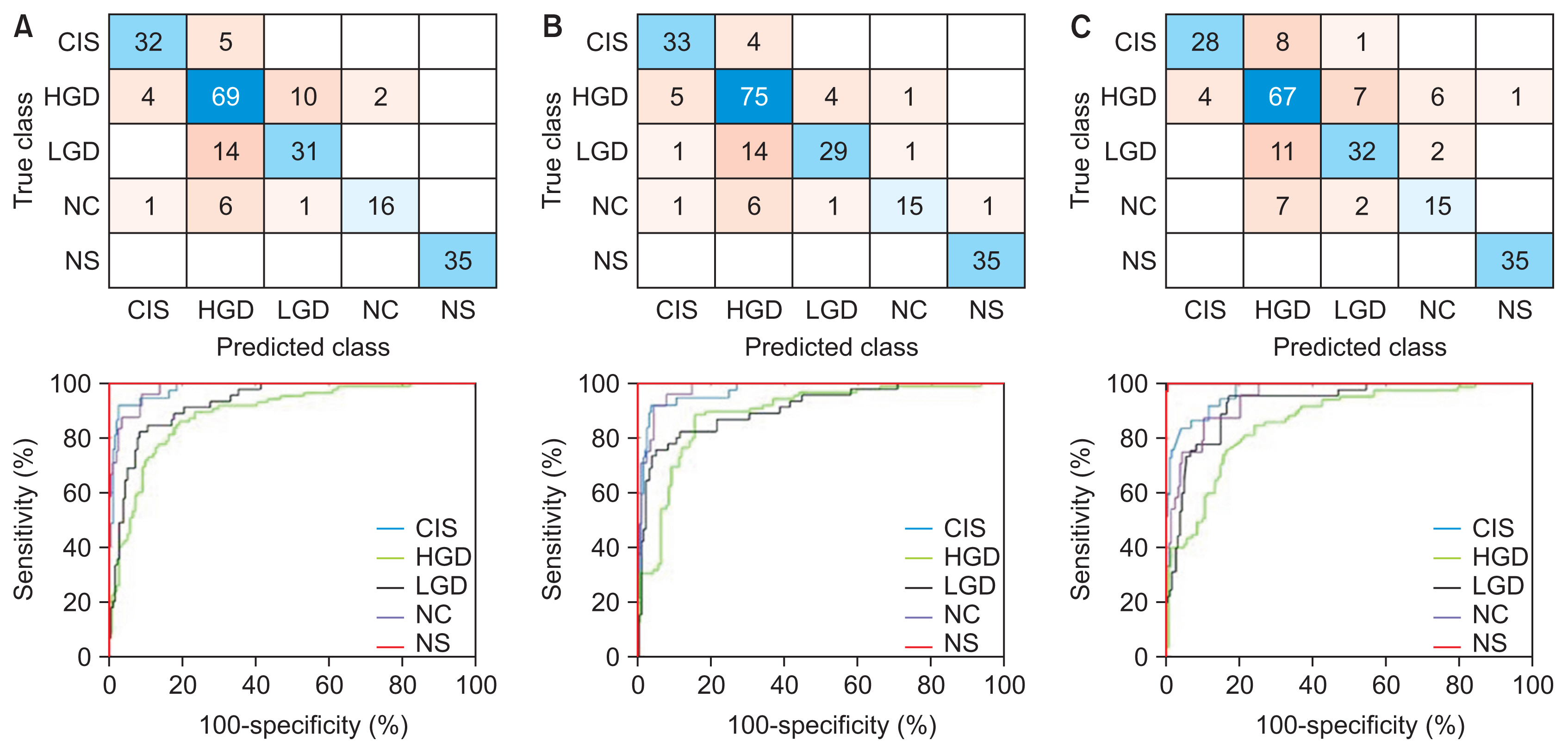
Figure 4
Classification sensitivity and sensitivity and area under the curve (AUC) values by cervical cell class of the best-performing AlexNet-SVM, VGG19-SVM, and ResNet50-SVM models. CIS: carcinoma, HGD: high grade, LGD: low grade dysplasia, NC: normal columnar, NS: normal squamous, SVM: support vector machine.
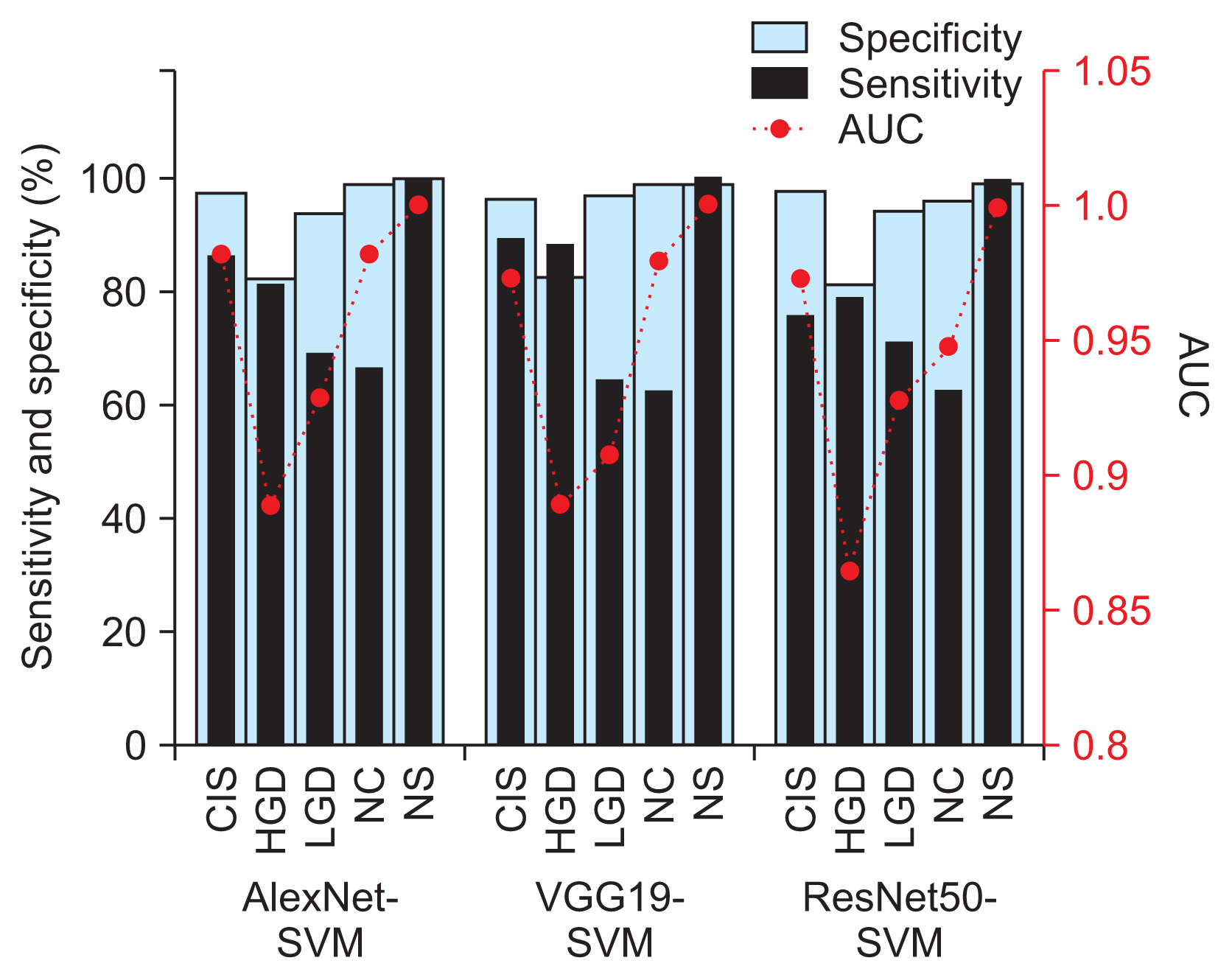
Table 1
Cervical cell dataset sizes and partitioning
Table 2
One versus one binary coding scheme
| Class | Binary learner | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | |
| CIS | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
| HGD | −1 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 |
| LGD | 0 | −1 | 0 | 0 | −1 | 0 | 0 | 1 | 1 | 0 |
| NC | 0 | 0 | −1 | 0 | 0 | −1 | 0 | −1 | 0 | 1 |
| NS | 0 | 0 | 0 | −1 | 0 | 0 | −1 | 0 | −1 | −1 |
Table 3
Performance of CNN-SVM models for cervical cell classification
Table 4
Performance comparison of classification methods
| Method | Experimental design | Average performance (%) | ||
|---|---|---|---|---|
| Accuracy | Specificity | Sensitivity | ||
| Fusion LeNet − SVM [18] | Single cell. Two-class problem | 99.30 | 99.40 | 98.90 |
| Inception-ResNet + snapshot ensemble [19] | Single cell. Seven-class problem | 65.56 | - | - |
| ConvNet + adaptive extraction [22] | Single cell. Two-class problem | 98.6 ± 0.3 | 99.0 ± 1.0 | 98.3 ± 0.7 |
| VGGNet + Mask R-CNN [20] | Single cell. Two-class and seven-class problem | 95.9 / 98.1 | 98.6 / 99.3 | 96.0 |
| Fusion ConvNet models − SVM [25] | Whole image. Four-class problem | 80.72 | - | - |
| Ensemble classifier (ResNet + GoogleNet) + majority voting [24] | Single cell. Four-class problem | 98 | 99 | 90 |
| AlexNet-SVM [16] | Single cell. Two-class problem | 99 | 97 | 99 |
| Current study | ||||
| AlexNet-SVM | Single cell. Five-class problem | 80.38 | 95.04 | 80.04 |
| VGG-SVM | Single cell. Five-class problem | 80.68 | 95.17 | 80.86 |
| ResNet-SVM | Single cell. Five-class problem | 77.00 | 93.98 | 75.92 |
References
1. Pelkofski E, Stine J, Wages NA, Gehrig PA, Kim KH, Cantrell LA. Cervical cancer in women aged 35 years and younger. Clin Ther 2016;38(3):459-66.


2. Yoruk S, Acikgoz A, Ergor G. Determination of knowledge levels, attitude and behaviors of female university students concerning cervical cancer, human papiloma virus and its vaccine. BMC Womens Health 2016;16:51.



3. Watson M, Benard V, King J, Crawford A, Saraiya M. National assessment of HPV and Pap tests: changes in cervical cancer screening, National Health Interview Survey. Prev Med 2017;100:243-7.



4. MacLaughlin KL, Jacobson RM, Radecki Breitkopf C, Wilson PM, Jacobson DJ, Fan C, et al. Trends over time in pap and pap-HPV cotesting for cervical cancer screening. J Womens Health (Larchmt) 2019;28(2):244-9.



5. Akbarzadeh MA, Hosseini MS. Implications for cancer care in Iran during COVID-19 pandemic. Radiother Oncol 2020;148:211-2.



6. Tsang-Wright F, Tasoulis MK, Roche N, MacNeill F. Breast cancer surgery after the COVID-19 pandemic. Future Oncol 2020;16(33):2687-90.


7. Price GJ, McCluggage WG, Morrison ML, McClean G, Venkatraman L, Diamond J, et al. Computerized diagnostic decision support system for the classification of preinvasive cervical squamous lesions. Hum Pathol 2003;34(11):1193-203.


8. Garrison A, Fischer AH, Karam AR, Leary A, Pieters RS. Cervical cancer. In: Pieters R, Rosenfeld J, Chen A, editors. Cancer concepts: a guidebook for the non-oncologist. Worcester (MA): University of Massachusetts Medical School; 2015.

9. William W, Ware A, Basaza-Ejiri AH, Obungoloch J. A review of image analysis and machine learning techniques for automated cervical cancer screening from pap-smear images. Comput Methods Programs Biomed 2018;164:15-22.


10. Selvathi D, Sharmila WR, Sankari PS. Advanced computational intelligence techniques based computer aided diagnosis system for cervical cancer detection using pap smear images. In: Dey N, Ashour AS, Borra S, editors. Classification in BioApps. Cham, Switzerland: Springer; 2008. p. 295-322.

11. Win KY, Choomchuay S, Hamamoto K, Raveesunthornkiat M, Rangsirattanakul L, Pongsawat S. Computer aided diagnosis system for detection of cancer cells on cytological pleural effusion images. Biomed Res Int 2018;2018:6456724.



12. Fekri-Ershad S. Pap smear classification using combination of global significant value, texture statistical features and time series features. Multimed Tools Appl 2019;78(22):31121-36.

13. Cervantes J, Garcia-Lamont F, Rodríguez-Mazahua L, Lopez A. A comprehensive survey on support vector machine classification: applications, challenges and trends. Neurocomputing 2020;408:189-215.

14. Chauhan VK, Dahiya K, Sharma A. Problem formulations and solvers in linear SVM: a review. Artif Intell Rev 2019;52(2):803-55.

15. Aoyagi K, Wang H, Sudo H, Chiba A. Simple method to construct process maps for additive manufacturing using a support vector machine. Addit Manuf 2019;27:353-62.

16. Taha B, Dias J, Werghi N. Classification of cervicalcancer using pap-smear images: a convolutional neural network approach. In: Valdes Hernandez M, Gonzalez-Castro V, editors. Medical image understanding and analysis. Cham, Switzerland: Springer; 2017. p. 261-72.

17. Khamparia A, Gupta D, de Albuquerque VH, Sangaiah AK, Jhaveri RH. Internet of health things-driven deep learning system for detection and classification of cervical cells using transfer learning. J Supercomput 2020;76:8590-608.

18. Jia AD, Li BZ, Zhang CC. Detection of cervical cancer cells based on strong feature CNN-SVM network. Neurocomputing 2020;411:112-27.

19. Chen W, Li X, Gao L, Shen W. Improving computer-aided cervical cells classification using transfer learning based snapshot ensemble. Appl Sci 2020;10(20):7292.

20. Kurnianingsih , Allehaibi KH, Nugroho LE, Lazuardi L, Prabuwono AS, Mantoro T. Segmentation and classification of cervical cells using deep learning. IEEE Access 2019;7:116925-41.

21. Ghoneim A, Muhammad G, Hossain MS. Cervical cancer classification using convolutional neural networks and extreme learning machines. Future Gener Comput Syst 2020;102:643-49.

22. Zhang L, Le Lu, Nogues I, Summers RM, Liu S, Yao J. DeepPap: deep convolutional networks for cervical cell classification. IEEE J Biomed Health Inform 2017;21(6):1633-43.


23. Wang P, Wang J, Li Y, Li L, Zhang H. Adaptive pruning of transfer learned deep convolutional neural network for classification of cervical pap smear images. IEEE Access 2020;8:50674-83.

24. Hussain E, Mahanta LB, Das CR, Talukdar RK. A comprehensive study on the multi-class cervical cancer diagnostic prediction on pap smear images using a fusion-based decision from ensemble deep convolutional neural network. Tissue Cell 2020;65:101347.


25. AlMubarak HA, Stanley J, Guo P, Long R, Antani S, Thoma G, et al. A hybrid deep learning and handcrafted feature approach for cervical cancer digital histology image classification. Int J Healthc Inf Syst Inform 2019;14(2):66-87.

26. Johnson DB, Rowlands CJ. Diagnosis and treatment of cervical intraepithelial neoplasia in general practice. BMJ 1989;299(6707):1083-6.



27. Mutombo AB, Simoens C, Tozin R, Bogers J, Van Geertruyden JP, Jacquemyn Y. Efficacy of commercially available biological agents for the topical treatment of cervical intraepithelial neoplasia: a systematic review. Syst Rev 2019;8(1):132.



28. Smith CM, Watson DI, Michael MZ, Hussey DJ. MicroRNAs, development of Barrett’s esophagus, and progression to esophageal adenocarcinoma. World J Gastroenterol 2010;16(5):531-7.



- TOOLS
-
METRICS

-
- 0 Crossref
- 3 Scopus
- 3,203 View
- 111 Download
- Related articles in Healthc Inform Res






